Levitra enthält Vardenafil, das eine kürzere Wirkdauer als Tadalafil hat, dafür aber schnell einsetzt. Männer, die diskret bestellen möchten, suchen häufig nach levitra kaufen ohne rezept. Dabei spielt die rechtliche Lage in der Schweiz eine wichtige Rolle.
Ibch.poznan.pl
THE JOURNAL OF BIOLOGICAL CHEMISTRY
Vol. 278, No. 48, Issue of November 28, pp. 47987–47996, 2003
2003 by The American Society for Biochemistry and Molecular Biology, Inc.
Printed in U.S.A.
Structural Rearrangements of the 10 –23 DNAzyme to 
3 Integrin
Subunit mRNA Induced by Cations and Their Relations to the
Catalytic Activity*
Received for publication, January 16, 2003, and in revised form, July 22, 2003
Published, JBC Papers in Press, September 2, 2003, DOI 10.1074/jbc.M300504200
Marcin Cieslak‡, Jacek Szymanski§, Ryszard W. Adamiak¶, and Czeslaw S. Cierniewski§储
**
From the Centers for ‡
Molecular and Macromolecular Studies and 储
Medical Biology and Microbiology, Polish Academy ofSciences, §
Department of Molecular and Medical Biophysics, Medical University of Lodz, 92-215 Lodz, Poland and¶
Institute of Bioorganic Chemistry, Polish Academy of Sciences, 61-704 Poznan, Poland
The intracellular ability of the "10 –23" DNAzyme to
biological processes such as tumor angiogenesis. For example,
efficiently inhibit expression of targeted proteins has
DNAzymes to 1 and 3 mRNA reduced expression of targeted
been evidenced by in vitro and in vivo studies. However,
integrin subunits in endothelial cells and blocked proliferation,
standard conditions for kinetic measurements of the
migration, and network formation in a fibrin and Matrigel™
DNAzyme catalytic activity in vitro include 25 mM Mg2ⴙ
,
matrix (2). In a cell culture system, a 10 –23 deoxyribozyme
a concentration that is very unlikely to be achieved
designed against 12-lipoxygenase mRNA specifically down-reg-
intracellularly. To study this discrepancy, we analyzed
ulated expression of this protein and its metabolites, which are
the folding transitions of the 10 –23 DNAzyme induced
known to play a crucial role in tumor angiogenesis (3). Simi-
by Mg2ⴙ
. For this purpose, spectroscopic analyzes such
larly, the DNAzyme to VEGFR2 mRNA cleaved its substrate
as fluorescence resonance energy transfer, fluorescence
efficiently and inhibited the proliferation of endothelial cells
anisotropy, circular dichroism, and surface plasmon
with a concomitant reduction of VEGFR2 mRNA and blocked
resonance measurements were performed. The global
geometry of the DNAzyme in the absence of added Mg2ⴙ
tumor growth
in vivo (4).
seems to be essentially extended, has no catalytic activ-
The origins of the DNAzyme catalytic activity are not yet
ity, and shows a very low binding affinity to its RNA
fully understood, but the observed rate enhancements probably
substrate. The folding of the DNAzyme induced by bind-
are generated by a number of factors, including metal ion and
ing of Mg2ⴙ
may occur in several distinct stages. The
nucleobase catalysis and local stereochemical effects. The
first stage, observed at 0.5 mM Mg2ⴙ
, corresponds to the
10 –23 DNAzyme has been developed using an
in vitro selection
formation of a compact structure with limited binding
strategy on the basis of its ability to cleave RNA in the presence
properties and without catalytic activity. Then, at 5 mM
of Mg2⫹ (1). It has a catalytic domain of 15 highly conserved
Mg2ⴙ
, flanking arms are projected at right position and
deoxyribonucleotides flanked by two substrate-recognition do-
angles to bind RNA. In such a state, DNAzyme shows
mains and can cleave effectively between any unpaired purine
substantial binding to its substrate and significant cat-
and pyrimidine of mRNA transcripts. Like many other en-
alytic activity. Finally, the transition occurring at 15 mM
zymes catalyzing phosphoryl-transfer reactions, it is recog-
Mg2ⴙ
leads to the formation of the catalytic domain, and
nized as a metalloenzyme requiring divalent metal, preferen-
DNAzyme shows high binding affinity toward substrate
tially Mg2⫹ ions, for catalytic activity. Divalent cations play a
and efficient catalytic activity. Under conditions simu-
crucial role in these mechanisms, as evidenced by a number of
lating intracellular conditions, the DNAzyme was only
partially folded, did not bind to its substrate, and
observations. For example, addition of La3⫹ to the Mg2⫹-back-
showed only residual catalytic activity, suggesting that
ground reaction mixture inhibited the DNAzyme-catalyzed re-
it may be inactive in the transfected cells and behave
actions, suggesting the replacement of catalytically and/or
like antisense oligodeoxynucleotide.
structurally important Mg2⫹ by La3⫹ (5). The function of diva-lent metal cations in DNAzyme activity is very complex andincludes (i) stabilization of the transition state of reaction. The
The typical DNAzyme,1 known as the "10 –23" model, has
divalent metal cation dependence of the enzyme was described
tremendous potential in gene suppression for both target vali-
as being the evidence supporting a chemical mechanism involv-
dation and therapeutic applications (1). It is capable of cleaving
ing metal-assisted deprotonation of the 2⬘-hydroxyl located ad-
single-stranded RNA at specific sites under simulated physio-
jacent to the cleavage site (6). It also includes (ii) neutralization
logical conditions and can be used to control even complex
of negative charges of phosphate groups, thus facilitating DNA-RNA interactions. High resolution x-ray crystal structuresof Mg2⫹ and Ca2⫹ salts of the model B-DNA decamers
* This work was supported by Grant Z-KBN 004/PO4/98 from the
CCAACGTTGG and CCAGCGCTGG revealed sequence-spe-
Polish Committee for Scientific Research. The costs of publication ofthis article were defrayed in part by the payment of page charges. This
cific binding of Mg2⫹ and Ca2⫹ to the major and minor grooves
article must therefore be hereby marked "
advertisement" in accordance
of DNA, as well as nonspecific binding to backbone phosphate
with 18 U.S.C. Section 1734 solely to indicate this fact.
oxygen atoms. This accounts for the neutralization of between
** To whom correspondence should be addressed: Dept. of Molecular
50 and 100% of the negative charges of phosphate groups (7).
and Medical Biophysics, Medical University of Lodz, 6/8 MazowieckaStreet, 92–215 Lodz, Poland. Tel.: 48-42-6783393; Fax: 48-42-6789433;
(iii) Some of these bound cations may also play a purely struc-
tural role by inducing proper folding of the DNAzyme molecule,
1 The abbreviations used are: DNAzyme, DNA enzyme suitable for
thus helping to organize the enzyme into its active conforma-
the sequence-specific cleavage of RNA; FRET, fluorescence resonance
tion. There are several reports showing that Mg2⫹ helps to
energy transfer; HPLC, high pressure liquid chromatography; HUVEC,human umbilical vein endothelial cell.
stabilize different types of double-stranded DNA structures (8,
This paper is available on line at http://www.jbc.org
Structural Transitions Induced in the DNAzyme by Mg2⫹
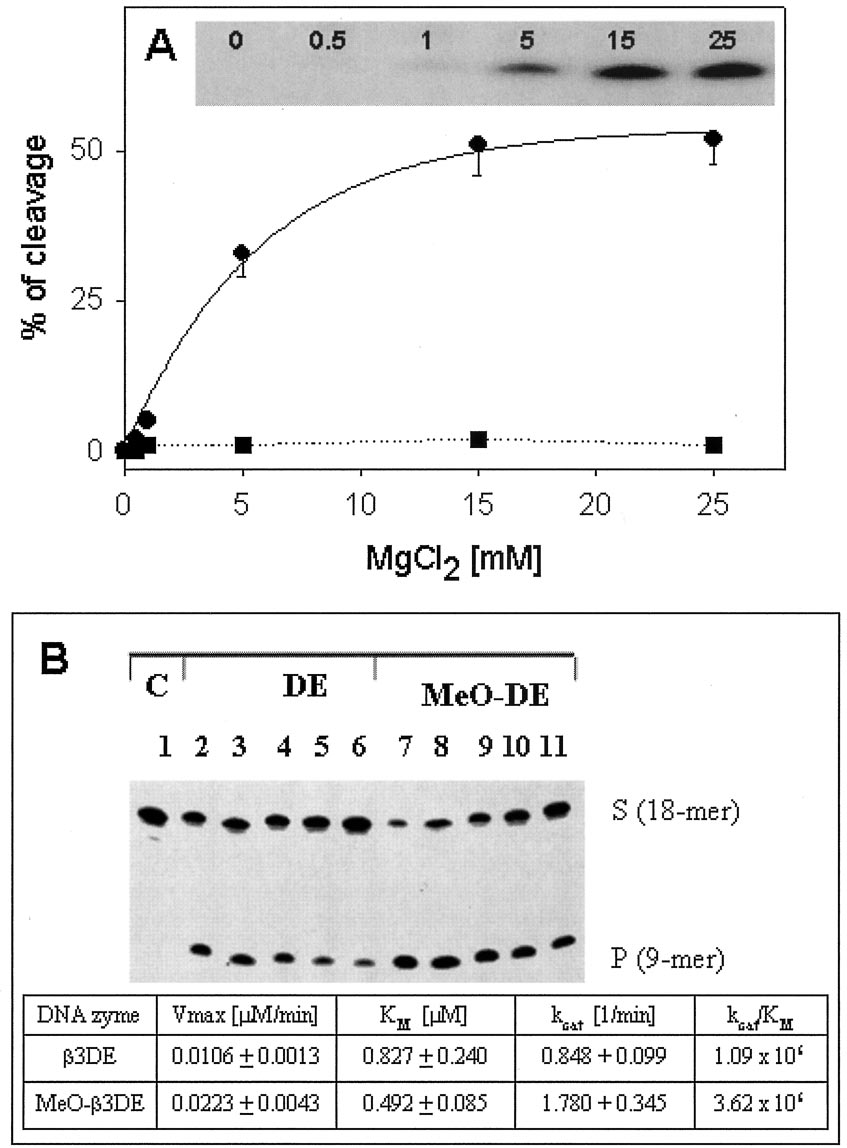
FIG. 1. Cleavage of 3 integrin subunit mRNA substrate by DNAzymes in vitro. A, the effect of Mg2⫹ concentration on enzymatic activity
of DNAzymes. MeO-3DE
(–) and its mutant MeO-3DE
(f–f) were incubated with the RNA substrate (molar ratio, 1:80) for 10 min in
the presence of increasing concentrations of Mg2⫹ ranging from 0 to 25 mM. The cleavage reaction was stopped by the addition of 0.5 M EDTA, andthe products were separated by electrophoresis in 20% polyacrylamide gels under denaturing conditions. Relative amounts of cleavage products(% of cleavage) are plotted versus Mg2⫹ concentration (MgCl [m
M]). Inset, autoradiogram of the gel showing the cleavage product obtained at
different Mg2⫹ concentrations (0 –25 mM). B, catalytic activity of 3DE and MeO-3DE. In these experiments, aliquots of the 32P-labeled mRNAsubstrate were incubated with DNAzymes. 3DE and MeO-3DE were used at a molar ratio (substrate:enzyme) ranging from 5:1 to 80:1 for upto 60 min at 37 °C. The cleavage products obtained after a 10-min incubation of 32P-labeled mRNA substrate are shown alone (lane 1) or withDNAzymes (3DE, MeO-3DE) mixed at the ratio 5:1 (lanes 2 and 7), 10:1 (lanes 3 and 8), 20:1 (lanes 4 and 9), 40:1 (lanes 5 and 10), and 80:1 (lanes6 and 11). DNAzymes were used at the concentration of 0.025 M. Reactions were carried out in 50 mM Tris, pH 8.0, containing 15 mM MgCl , 0.01%
SDS. Amounts of the product were evaluated by a PhosphorImager (Amersham Biosciences) and used to calculate kinetic parameters. They weredetermined in multiple turnover reactions and represent a mean of three independent experiments.
9) and can induce bending or enhance curvature in DNA (10).
with self-complementary ends (11). These structural effects of
Furthermore, Mg2⫹ and other divalent cations enhance end-to-
cations may be even more profound in a single-stranded and
end DNA interactions, particularly in the case of fragments
flexible DNAzyme molecule.
Structural Transitions Induced in the DNAzyme by Mg2⫹
Mg2⫹-dependent cleavage has special relevance to biology
in vivo (12). However, at the present time, it is hard to
because it is compatible with intracellular conditions, raising
explain their intracellular catalytic activity, keeping in mind
the possibility that DNA enzymes might be made to operate
the catalytic dependence upon high concentrations of Mg2⫹,which is unlikely to be achieved in cytoplasm. To address thequestion of their intracellular catalytic activity, we at-tempted to correlate changes in the catalytic activity andconfiguration of the DNAzyme induced by gradually boundMg2⫹. To characterize structural changes in the DNAzyme,we performed fluorescence resonance energy transfer (FRET)analysis, which allowed us to monitor general folding of themolecule based on the measurements of distances betweenfluorophores linked to 5⬘ and 3⬘ side bases, and surface plas-mon resonance analysis of the DNAzyme binding to its RNAsubstrate. Structural changes induced by cations inDNAzymes were also monitored by circular dichroism andfluorescence anisotropy analysis.
EXPERIMENTAL PROCEDURES
Synthesis of DNAzyme to 3 mRNA—DNAzyme was chemically syn-
thesized on a solid support using an ABI-394 DNA synthesizer, asdescribed previously (13). This particular DNA sequence (5⬘-GAGTCCCATAg g c t a g c t a c a a c g a AAGACTTGAG-3⬘)
was used previously to analyze the enzymatic activity, specificity, exo-nuclease resistance, and ability to inhibit expression of 3 integrins inendothelial cells (2). For BIAcore experiments, the inactive DNAzyme,
FIG. 2. Biological activity of Fluo-MeO-3DE-Rhod. A,
, with a single substitution (G6 3 A) in the reactive loop and
DNAzyme activity of the fluorophore-labeled construct identical to that
antisense oligodeoxynucleotide 3(1245–1265) containing both flank-
used for the FRET analysis. The cleavage activity was examined after
ing arms of the 3DE (5⬘-GAGTCCCATACAAGACTTGAG-3⬘) were
incubation of Fluo-MeO-3DE-Rhod with a 20-fold excess of 5⬘-32P-
synthesized. Two analogues of 3DE
and 3(1245–1265) were
labeled RNA substrate in the presence of 15 mM MgCl at 37 °C. A
produced as well, which contained the modified oligonucleotides such
sample was removed at different time points (lanes 1-5). B, the fluores-
as phosphorothioates or 2⬘-O-methyl-substituted residues introduced
cence image of endothelial cells treated with Fluo-MeO-3DE-Rhod.
at both 5⬘ and 3⬘ sides. Hence, S-3DE
and S-3(1245–1265) have
Endothelial cells, exposed to 0.5 M of the fluorophore-labeled construct
two phosphorothioate substitutions, whereas MeO-3DE
for 24 h at 37 °C and processed as described under ‘‘Experimental
MeO-3(1245–1265) contain two 2⬘-O-methyl-substituted residues at
Procedures,'' were analyzed by confocal fluorescence microscopy. The
both their 5⬘ and 3⬘ sides, respectively. An additional mutated
punctate fluorescence distribution within the cytoplasm was detected
DNAzyme (MeO-3DE
) with a downsized catalytic loop of a
by monitoring both fluorescein and rhodamine attached to DNAzyme.
C, the reduced expression of 3 mRNA in HUVECs treated with MeO-
TAg g c t a c a a c g a AAGACTTGAG-3⬘) was synthesized
3DE when compared with unchanged expression in untreated cells. 3
and used as a control. DNAzymes with the 11-mer catalytic loop were
mRNA was evaluated by relative quantitative reverse transcriptase-PCR using glyceraldehyde-3-phosphate dehydrogenase mRNA as an
described to be Ca2⫹-dependent deoxyribozymes and showed signifi-
intrinsic control.
cantly reduced binding affinities and catalytic activities in the pres-
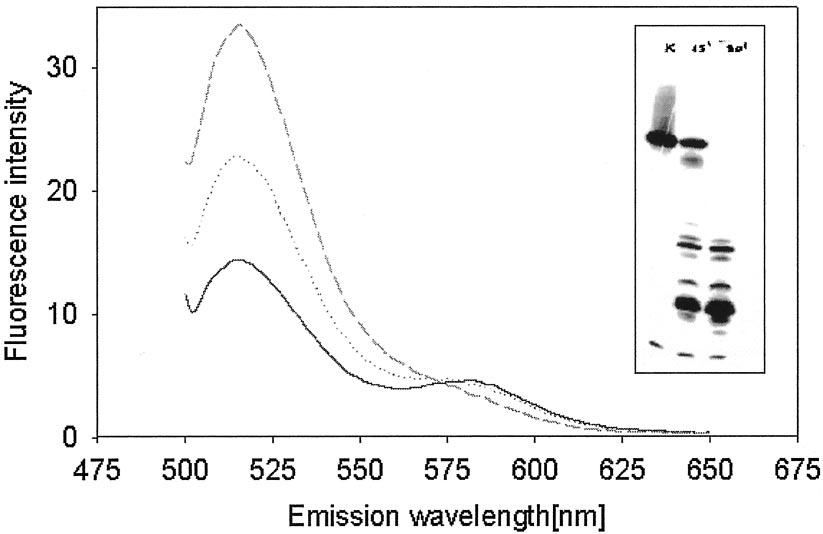
FIG. 3. Fluorescence resonance energy transfer between fluorescein and rhodamine attached to MeO-3DE at 3ⴕ and 5ⴕ ends,
respectively. Fluo-MeO-3DE-Rhod (40 nM) had two peaks at 520 nm and 580 nm upon excitation at 494 nm. Inset, autoradiogram of a gel after
electrophoresis of the Fluo-MeO-3DE-Rhod digested with endonuclease from S. marcescens. Degradation of the fluorophore-labeled DNAzyme
resulted in a significant increase in fluorescence intensity at 520 nm and a disappearance of the 580 nm maximum.
Structural Transitions Induced in the DNAzyme by Mg2⫹
ence of Mg2⫹ (14). All deoxyoligonucleotides were purified by HPLC
matically corrected for lamp intensity variations. Buffers were de-
and ion exchange chromatography (to 98%), and their purity was
gassed by bubbling nitrogen to prevent quenching of fluorescence by
checked by PAGE under denaturing conditions.
dissolved oxygen. The fluorescence emission signals were stable to
The doubly labeled 3 DNAzymes with the 15-mer and 11-mer cat-
photobleaching under the experimental conditions of measurement.
alytic loops were synthesized by the solid-state phosphoramidite ap-
The apparent interchromophore separation, R, the distance separat-
proach on an ABI 392 synthesizer, starting from fluorescein CPG sup-
ing the energy donor and acceptor was calculated by the Forster
port (CPG, Inc.). The 2⬘-O-Me RNA cyanoethyl phosphoramidites (Glen
equation: r ⫽ R (1/E ⫺ 1)1/6, where E is the efficiency of energy
Research) were used for introduction of 2⬘-O-Me nucleotide units flank-
transfer from donor to acceptor. E ⫽ 1 ⫺ F
ing the sequence from the 3⬘ and 5⬘ ends. The synthesis was terminated
distance for 50% transfer efficiency E. F
with the addition of rhodamine cyanoethyl phosphoramidite (CPG,
cence intensities of the donor in the presence and absence of the
Inc.). Oligonucleotides were cleaved from the solid support and depro-
acceptor, respectively. F
were measured at 520 nm, the
tected by brief (4 h at 55 °C) treatment with 30% ammonia. Pure
emission maximum for fluorescein.
product was isolated by preparative HPLC on a Hamilton PRP1 col-
The fluorescence anisotropy of the fluorescein and rhodamine probes
umn. The peak fractions were evaporated to dryness, redissolved in
attached to the DNAzyme (Fluo-MeO-3DE
water, and then ethanol-precipitated.
3DE -Rhod) was monitored in an LS-50 spectrofluorometer
The synthetic, biotinylated (at the 3⬘ end) 21-mer RNA (biotin-CU-
(PerkinElmer Life Sciences) equipped with an automatic anisotropy
CAGGGUAUGUUCUGAACUC) used in BIAcore experiments was pur-
measuring device. The anisotropy r is defined as
chased from Bionovo (Poland). Its sequence corresponded to the 1245–1265 fragment of 3 mRNA. After synthesis, the product was purified
r ⫽ (I
G ⫻ IVH) / (IVV
2 ⫻ G ⫻ IVH)
by HPLC and its purity was checked by PAGE.
where I is fluorescence intensity. The first and second indices refer to
Preparation of Target RNA Substrates and Kinetic Analysis—Ali-
the orientation of excitation and emission polarizers, respectively. G is
quots of RNA substrates (20 l, 5 M) dissolved in a T4 polynucleotide
the correction factor. The cell holder was thermostated at 21 °C.
kinase buffer were mixed with [␥-32P]ATP (2 l, 20 Ci) and T4 polynu-
Analysis of Fluorescence Data—Efficiencies of energy transfer were
cleotide kinase (3 units). Reaction was carried out for an h at 37 °C. All
determined from enhancement acceptor fluorescence (17, 18). The
reported kinetic values were determined in multiple turnover reactions.
emission at a given wavelength (v1) of a double-labeled sample ex-
values were determined from the y intercept and slope,
cited primarily at the donor wavelength (v⬘) contains emission from
respectively, of the best-fit line to a Lineweaver-Burke plot of 1/V versus
the donor, emission from directly excited acceptor, and emission from
1/[S]. Reactions (10 min at 37 °C; total volume ⫽ 20 l) were carried out
acceptor excited by energy transfer from the donor, i.e.
in 50 mM Tris, pH 8.0, containing 15 mM MgCl , 0.01% SDS, DNAzyme
(0.0125 M) with the radiolabeled RNA substrate used in a wide range
F(v1,v⬘) ⫽ [S]䡠⑀D(v⬘)䡠D(v1)䡠d⫹䡠{(1 ⫺ EFRET)䡠a⫹ ⫹ a⫺
of concentrations. The cleavage reaction was stopped by the addition of5 l of 0.5 M EDTA, and the products were separated by electrophoresis
⫹ ⑀A(v⬘)䡠
A(v1)䡠a⫹ ⫹ ⑀D(v⬘)䡠A(v1)䡠EFRET d⫹ ⫹ a⫹}
in 20% polyacrylamide gels under denaturing conditions. Amounts ofthe product were evaluated by use of a PhosphorImager (Amersham
⫽ FD(v1,v⬘) ⫹ FA(v1,v⬘) (Eq. 2)
Cell Culture—Human umbilical vein endothelial cells (HUVEC)
where [S] is the concentration of DNAzyme, d⫹ and a⫹ are the molar
were isolated from freshly collected umbilical cords by collagenase
fraction of DNAzyme molecules labeled with donor and acceptor respec-
treatment (15, 16). Cells were grown in gelatin-coated 75-cm2 tissue
tively, and a⫺ is the molar fraction of DNAzyme molecules unlabeled
culture flasks and were maintained at confluence in RPMI 1640
with acceptor. Superscripts D and A refer to donor and acceptor, re-
medium supplemented with streptomycin (100 g/ml), penicillin (100
spectively. ⑀D(v⬘) and ⑀A(v⬘) are the molar absorption coefficients of
units/ml), fungizone (2.5 mg/ml), heparin (90 g/ml),
donor and acceptor, respectively, and (v ) and (v ) are the fluores-
cent quantum yields of donor and acceptor, respectively. Thus the
M), sodium bicarbonate (2 mg/ml), 20% fetal bovine serum, and
epidermal growth factor (40 ng/ml) at 37 °C in a humidified 5%
spectrum contains the components due to donor emission [FD(v ,v⬘), i.e.
atmosphere. Primary cultures were harvested at confluence
the first term containing (v )] and those due to acceptor emission
with trypsin/EDTA and transferred into gelatin-coated dishes. For
[FA(v ,v⬘), i.e. the latter two terms containing (v )]. The first stage of
the experiments, confluent cultures were used at the second
the analysis involves subtraction of the spectrum of DNA labeled only
with donor, leaving just the acceptor components, i.e. FA(v ,v⬘). The
For microscopic examination, cells were plated at a density of 5 ⫻ 104
pure acceptor spectrum thus derived is normalized to one from the same
cells/well on Thermanox cover-slips in 8-well tissue culture chamber
sample excited at a wavelength (v⬙) at which only the acceptor is
slides (NUNC) with detachable chambered upper structures. Before
excited, with emission at v2. We then obtain the acceptor ratio
performance of assays, the serum-containing medium was changed to a
serum-free medium (Opti-MEM). The cultures were gently rinsed three
1,v⬘) / FA(v2,v⬙)
times with the medium and preincubated with fluorophore-labeled
FRET d⫹䡠[⑀D(v⬘) / ⑀A(v⬙) ⫹ ⑀A(v⬘) / ⑀A(v⬙)]}䡠[A(v1) / A(v2)]
M) for 6 h in the presence of Lipofectin (5 g/ml).
After that time, the transfection mixture was replaced by normal se-
is directly proportional to (ratio) and can be easily calculated
rum-containing medium, and cells were grown for another 18 h. At-
because ⑀D(v⬘)/⑀A(v⬙) and ⑀A(v⬘)/⑀A(v⬙) are measured from absorption
tached, treated intact cells were maintained in a CO incubator at
spectra, and (v )/ (v ) is unity when v ⫽ v .
37 °C. Two control assays were carried out using either untreated cells
Analysis of Circular Dichroism—The circular dichroism spectra of
or cells exposed to 0.25 M fluorescein. After incubation, cells were
MeO-3DE (1 M), free or in the complex with the substrate (2 M), was
washed three times with phosphate-buffered saline, fixed with freshly
measured in a solution of 10 mM Tris-HCl, pH 7.5, containing increasing
prepared 3.5% paraformaldehyde for 15 min at room temperature,
concentrations of MgCl at 21 °C. In control experiments, 0.1
washed three times with phosphate-buffered saline, mounted in 2.5%
NaCl were used. Before measurement, the complex was allowed to form
DABCO™ in glycerol, and processed for microscopy.
by heating the solution at 95 °C for 2.5 min followed by gradual cooling.
Fluorescence Spectroscopy—Fluorescence emission spectra were
Measurements were made in a quartz cuvette (5-mm path length) with
measured on an LS-50 spectrofluorometer (PerkinElmer Life Sciences),
a CD spectrometer (model CD6, Jobin Yvon) from 200 to 320 nm in
and spectra were corrected for lamp fluctuations and instrumental
triplicate. The spectra were obtained by smoothing the averaged spec-
variations. Polarization artifacts were avoided by setting excitation and
tra with a calculator.
emission polarizers to magic angle conditions (54.74°). All of the fluo-
Surface Plasmon Resonance—The kinetic parameters (association
rescence measurements were performed at the temperature of 23 ⫾
and dissociation rate constants, k
and k , respectively) and the af-
1 °C. Emission spectra, excitation spectra, and luminescence intensity
finity constant (K ) between DNAzyme and the mRNA substrate were
were recorded with 5-nm band passes for both the excitation and
measured by surface plasmon resonance using a BIAcoreX (Amersham
emission monochromators. A cut-off filter in the emission beam was
Biosciences). Briefly, avidin was covalently attached to carboxymethyl
used to eliminate second-order wavelength interference. The excita-
dextran chips (CM5, BIAcore) previously activated with N-hydroxysuc-
tion wavelengths used were 494 nm and 560 nm for fluorescein- and
cinimide and N-ethyl-N⬘-dimethylaminopropyl carbodiimide, according
rhodamine-conjugated constructs, respectively. Emission spectra
to the manufacturer's instructions. Experiments were performed at
were corrected for the blank contribution and for the instrument
37 °C using 50 mM Tris, pH 8.0, containing divalent cations at the
response and normalized to the DNA concentration in a quartz cell
indicated concentrations. 15 l of 5⬘ end-biotinylated RNA at 100 nM in
with a 1-cm path length. Excitation and emission spectra were auto-
50 mM Tris, pH 8.0, was injected at the flow rate of 5 l/min and
Structural Transitions Induced in the DNAzyme by Mg2⫹
consequently immobilized on the bound avidin to give a response of
same as those recently reported for another 10 –23 DNAzyme
⬃500 resonance units, an arbitrary unit specific for the BIAcore instru-
(21). Cellular transport of the Fluo-MeO-3DE
ment. The levels of immobilized RNA were within the low levels that
function of the external oligonucleotide concentration was non-
have to be used to ensure that the observed binding rate will be limitedby the reaction kinetics rather than by the mass transport effects of the
linear, being more efficient at concentrations below 2 M. The
injected DNAzyme (19). In typical experiments, DNAzyme flowed in
punctate fluorescence distribution observed even after 24 h of
two channels of the sensor; the first one contained the RNA substrate
exposure to the DNAzyme seems to suggest that endosomal
attached to avidin, and the second was without the RNA substrate. The
vesicles are the primary targets of the probes under study (Fig.
latter was used to correct SPR traces and remove the background
-Rhod could be detected intracellu-
binding between DNAzyme and the immobilized avidin on the dextran.
larly, both when emission of either fluorescein or rhodamine
The concentration of the injected DNAzyme was in the range 20 –200nM, and the flow rate was 5 l/min. The amount of ligand bound to
was measured and both fluorophores showed full colocaliza-
immobilized RNA substrate was monitored by measuring the variation
tion. Thus, the attached fluorophores did not influence the
of the surface plasmon resonance angle as a function of time. Results
enzymatic activity and biological properties of the DNAzyme,
were expressed in resonance units. In preliminary experiments, the
including the ability to interact with cellular components re-
data obtained for at least three different concentrations of DNAzymes
sponsible for its transport. Transfection of endothelial cells
were fitted to several models; the best fits (2 ⫽ 1.4) were obtained byassuming a one-to-one interaction. Then, the association rate constant,
with the DNAzyme (5 M) efficiently reduced expression of 3
k , and dissociation rate constant, k , were determined separately
integrin subunit measured at the level of 3 mRNA by reverse
from individual association and dissociation phases, respectively. The
transcriptase-PCR (Fig. 2C) and at the cellular surface by flow
overall affinity constant, K , was derived from k /k . The sensor chip
cytometry (not shown). Although MeO-3DE
was regenerated with three 10-l pulses of 12.5% formamide.
same cellular distribution as MeO-3DE
-Rhod, it did not
show any biological activity detectable at the level of 3 mRNA
or 3 expression at the cell surface. These data provide the
Enzymatic Characteristics of the DNAzyme to 3 Integrin
evidence that the 10 –23 DNAzyme has an intracellular cata-
Subunit—The 10 –23 DNAzyme used in these studies was de-
lytic or antisense activity, even at the much lower cation con-
signed to cleave 3 integrin subunit mRNA, and preliminary
centrations than those normally used in in vitro analysis.
characterization was done in our recent work (2). Standard
conditions for kinetic measurements of the catalytic activity of
Rhod were next used to measure energy transfer resulting from
DNAzyme in vitro include 25 mM MgCl
a dipolar coupling between the transition moments of the two
conditions, the 10 –23 DNAzyme shows optimal enzymatic ac-
fluorophores, fluorescein as the energy donor and rhodamine as
tivity, whereas its mutant MeO-3DE
with the shortened
the energy acceptor. When the fluorescence spectrum of one
catalytic loop is inactive (Fig. 1A). In this experiment, enzy-
fluorophore (the donor) overlaps with the excitation spectrum
matic reactions were performed in 50 mM Tris-HCl, pH 8.0,
of another fluorophore (the acceptor), the excitation of the
containing 0 –25 mM MgCl , under multiple turnover condi-
donor induces fluorescence of the acceptor, although its own
tions. The 32P-labeled mRNA substrate was mixed with the
fluorescence decreases. The extent of FRET is extremely sen-
DNAzyme 3DE in the molar ratio of 80:1 and incubated at
sitive to the distance between the donor and the acceptor, being
37 °C; aliquots were withdrawn after 10 min. The cleavage
inversely proportional to the sixth power of the distance. At-
reaction was stopped by the addition of 5 l of 0.5 M EDTA, and
tachment of rhodamine to Fluo-MeO-3DE
products were separated by electrophoresis in 20% polyacryl-
decrease in the fluorescence emission at 520 nm characteristic
amide gels under denaturing conditions. Amounts of the prod-
for fluorescein, and the fluorescence spectrum of the resulting
uct were evaluated by use of a PhosphorImager. Fig. 1B showsa composition of cleavage mixtures obtained after a 60-min
-Rhod, had two peaks at 520 nm
incubation of 32P-labeled mRNA substrates with 0.025
and 580 nm upon excitation at 494 nm (Fig. 3). The mutated
3DE added at a molar ratio ranging from 5:1 to 80:1. Each of
3DE, Fluo-MeO-3DE
-Rhod, showed the same fluores-
the DNAzymes, unmodified (3DE) and modified (MeO-3DE),
cence properties. These spectra clearly indicate that consider-
cleaved the substrate at the predicted site. Interestingly,
able energy from the excited fluorescein was transferred to
DNAzyme with the 2⬘-MeO residues showed a significantly
rhodamine, providing the evidence that both fluorophores are
higher enzymatic activity than 3DE, as evidenced by the
in close proximity. Cleavage of the Fluo-MeO-3DE
catalytic efficiency k
of 3.62 ⫻ 106 and 1.09 ⫻ 106 ⫺1
with endonuclease from Serratia marcescens resulted in a sig-
nificant increase of the fluorescence intensity at 520 nm, which
Study of Ion-induced Folding of the DNAzyme by Fluores-
approaches the level of fluorescein alone, indicating that both
cence Resonance Energy Transfer—To follow the folding tran-
fluorophores are separated to the distance enabling the energy
sitions of the DNAzyme induced upon binding of Mg2⫹ ions,
FRET was utilized. For this purpose, 3DE with the 15-mer
Variation in End-to-end Distances during the Ion-induced
and 11-mer catalytic loops was modified by attachment of the
Folding of the DNAzyme—According to the model for the fold-
donor and acceptor fluorophores, rhodamine and fluorescein, to
ing of the DNAzyme, the length of oligodeoxynucleotide should
the 5⬘ and 3⬘ ends of the flanking arms, respectively. Next, a
shorten over the full range of Mg2⫹ concentration. Experimen-
series of experiments were designed to analyze whether the
tally, we find that E
increases rapidly upon the addition of
fluorophores attached to the terminal bases affect the ability of
cations to Fluo-MeO-3DE
-Rhod and reaches a plateau
the oligodeoxynucleotide construct to function as an active
value by 5 mM Mg2⫹. Assuming a Forster critical distance (R )
DNAzyme species. As seen in Fig. 2A, incubation of the Fluo-
of 5.5 nm for donor-fluorescein (22), the distance between donor
-Rhod with the 32P-labeled substrate in the pres-
(fluorescein) and acceptor (rhodamine) in the absence of Mg2⫹
ence of 25 mM Mg2⫹ under multiple-turnover conditions at
was calculated to be 7.82 ⫹ 0.39 nm and was not significantly
37 °C leads to cleavage at the correct site. When such a con-
dependent upon the DNAzyme concentration. The distance
struct was incubated with endothelial cells, it remained resist-
between two fluorophores shortens over this range from 7.82
ant to intracellular nucleases and, even after 24 h, was located
nm to 6.22 nm and further addition of Mg2⫹ ions does not
exclusively within the cytoplasm, particularly in the perinu-
essentially change it (Fig. 4A). At 25 mM Mg2⫹, the R value
clear organelles. The cellular uptake, intracellular distribu-
reaches 6.11 nm and is almost identical to that characteristic
tion, and stability of the Fluo-MeO-3DE
for the Fluo-MeO-3DE-Rhod in the complex with its mRNA
Structural Transitions Induced in the DNAzyme by Mg2⫹
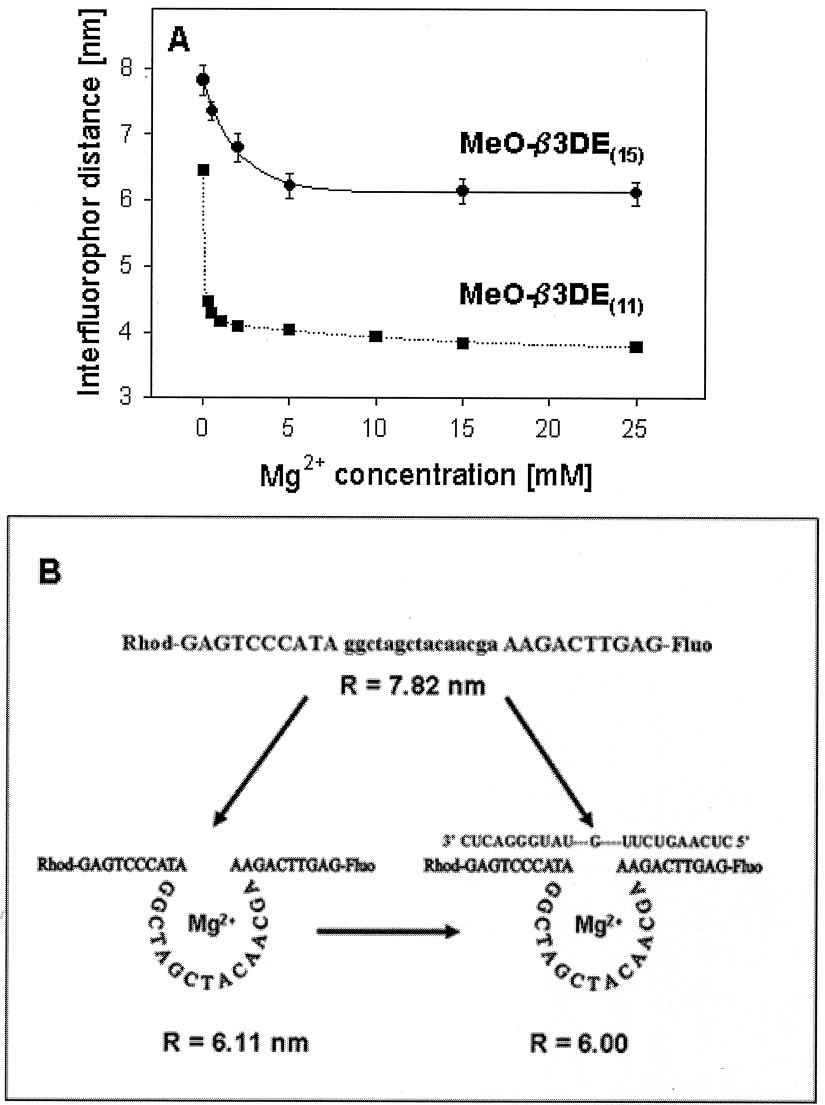
FIG. 4. FRET analysis for the MeO-3DE as a function of Mg2ⴙ concentration. A, calculated interfluorophore distances based on
measured efficiency of energy transfer presented as a function of MgCl concentration. The plot shows the variation in FRET efficiency as a
function of Mg2⫹ concentration up to 25 mM when Fluo-MeO-3DE
-Rhod (–) or Fluo-MeO-3DE
-Rhod (f–f) were tested. In accordance
with the model, the FRET efficiency is found to increase (indicating a reducing end-to-end distance) over this complete range. B, scheme with theexpected behavior of the free DNAzyme or complexed with its RNA substrate in the presence of Mg2⫹. The distances (R) were calculated based onFRET analysis.
substrate. Essentially the same changes in the interfluoro-
bozyme by means of high affinity binding to sites (Fig. 4B).
phore distance were induced in the Fluo-MeO-3DE-Rhod
To evaluate overall changes induced in the DNAzyme by
upon binding of other divalent cations, such as Ca2⫹ and Mn2⫹
cations, the fluorescence anisotropy of both fluorophores in
(Table I). Such folding of the DNAzyme does not result simply
Fluo-MeO-3DE-Rhod was measured in the presence of in-
from the neutralization of the polyanionic nature of the oligode-
creasing concentrations of Mg2⫹, Ca2⫹, Li⫹, Na⫹, and K⫹. The
oxynucleotide, because neither Na⫹ nor K⫹ added in place of
fluorescence anisotropy r reflects the local and global motions
Mg2⫹, even at 1 M, showed any effect. In the case of DNAzyme
of the fluorophore and is close to zero for a freely rotating
with the 11-mer loop, Fluo-MeO-3DE
fluorophore. The theoretical upper limit of 0.4 corresponds to a
matically increased upon the addition of 0.5 mM Mg2⫹, indicat-
totally non-rotating fluorophore (23). The fluorescein- and rho-
ing that these cations induce folding of the mutated deoxyri-
damine-labeled DNAzyme has a flexible single-stranded mole-
Structural Transitions Induced in the DNAzyme by Mg2⫹
cule characterized by a low r value ⫽ 0.016 and 0.037, as
or Ca2⫹ concentration reached 5 mM or 1 mM, respectively,
measured for fluorescein and rhodamine, respectively (Fig. 5).
indicating the increased condensation state of the molecule.
In both cases, the fluorescence anisotropy doubles when Mg2⫹
However, there was no change in the fluorescence anisotropywhen monovalent cations were used even at much higher con-
centrations (Fig. 5). These results are fully consistent with the
The estimated interfluorophore distance based on
proposed mechanism involving the divalent cation-induced
FRET analysis of the Fluo-MeO-3 DE-Rhod
folding of the DNAzyme and indicate that its molecule becomes
The fluorophore-labeled MeO-3DE (40 nM) was incubated with dif-
more compact upon Mg2⫹ binding.
ferent divalent cations used in the concentration range from 0 to 25 mM,and the interfluorophore distance R was evaluated based upon the
Complex Formation between DNAzyme and its RNA Sub-
Forster equation. Data represent a mean value of three separate deter-
strate—The conformational changes of the DNAzyme induced
by Mg2⫹ were next analyzed by CD spectroscopy (Fig. 6). CD
Interfluorophore distance [nM]
spectra of an ion-free DNAzyme/RNA complex were reported
earlier (24). We have been interested in examining how far CDspectroscopy (being sensitive to the structure helicity and fold)
could be applicable to probing binding of metal ions to single-
and double-stranded species of interest. As the reference, CD
spectra related to a regular complex formation (Fig. 6A), re-
flecting an antisense mechanism, were inspected first. The CD
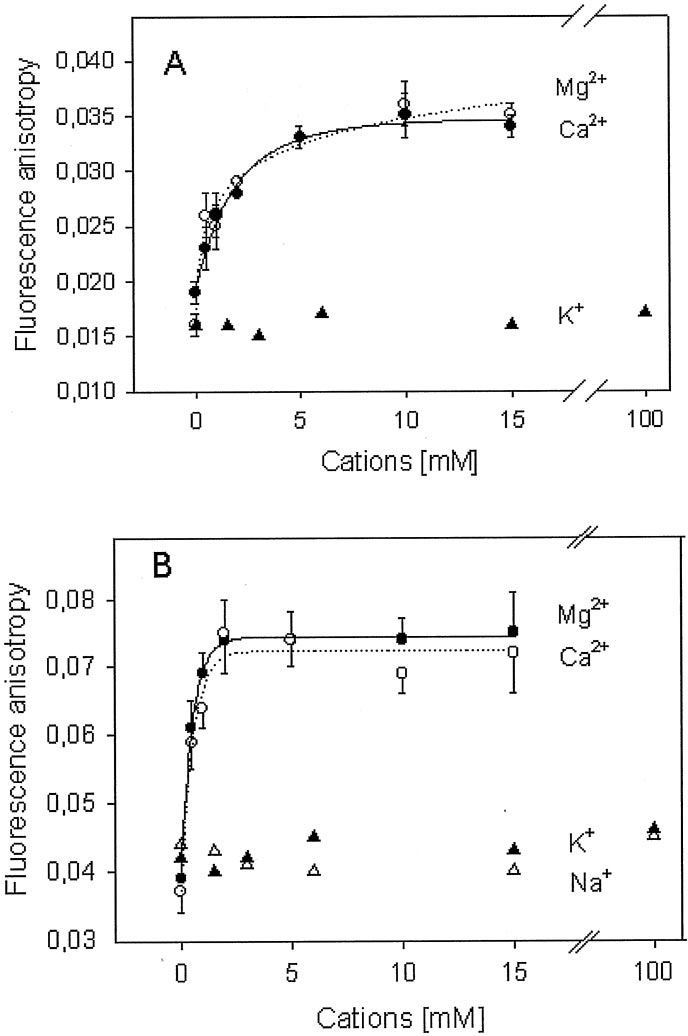
FIG. 5. Effect of Mg2ⴙ on the fluores-
cence anisotropy of Fluo-MeO-3DE-
Rhod. MgCl (–), CaCl (E–E), KCl
(Œ–Œ), and NaCl (‚–‚) were added step-wise to 0.5 M Fluo-MeO-3DE
and incubated for 5 min before measuringthe fluorescence anisotropy. To monitorthe fluorescence anisotropy of fluorescein(A) or rhodamine (B), the samples wereexcited at 494 nm or 560 nm and fluores-cence emission was measured at 520 nmor 580 nm, respectively.
Structural Transitions Induced in the DNAzyme by Mg2⫹
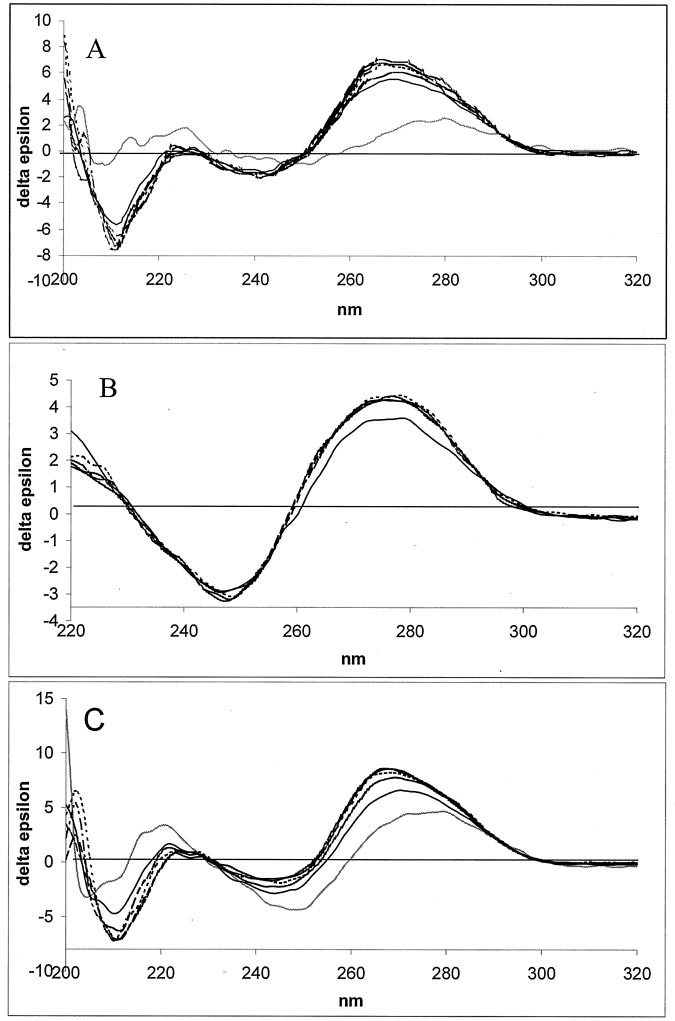
FIG. 6. Effect of Mg2ⴙ on CD spectra
of the free MeO-3DE and its com-
plex with the target RNA. CD spectra
of
MeO-3(1245–1265)/RNA complex (A),the single stranded MeO-3DE (B), andthe MeO-3DE/RNA complex (C) weretaken in the presence of Mg2⫹ concentra-tions ranging from 0 to 25 mM. However,because there was no change when con-centrations higher than 5 mM Mg2⫹ wereused, those spectra were deleted for clar-ity of the plot. Spectra of the single-stranded antisense oligodeoxynucleotideMeO-3(1245–1265) and MeO-3DE areshown as a gray line in A and C, respec-tively. Spectra were taken at the follow-ing Mg2⫹ concentrations: 0 mM (—), 0.5mM (----), 1.0 mM (䡠 䡠 䡠 䡠), 2.0 mM (-䡠 -䡠 -䡠 -䡠),and 5.0 mM (-䡠 䡠 -䡠 䡠 -䡠).
spectrum of a non-enzymatic, single 21-mer DNA strand is
strand of MeO-3DE, most probably close to that predicted by
irregular and of low magnitude. Addition of a complementary
the DNA-folding algorithm (26). An addition of Mg2⫹ at the
RNA strand resulted in raising a regular positive Cotton effect
initial concentration level (0.5 mM) led to a formation of a more
at 269 nm, i.e. a region typical of the DNA/RNA hybrids (25).
regular and higher amplitude positive band at 276 nm. Further
Influence of an increased Mg2⫹ concentration on a double hel-
Mg2⫹ additions (up to 25 mM) had no practical effect on the
ical structure is not strong but clearly visible, as indicated by
DNAzyme strand spectra. The binding of the Mg2⫹-free MeO-
the increase in amplitude of the Cotton effect. An initial addi-
3DE strand to the target RNA leads to the formation of a new
tion of Mg2⫹ (0.5 mM) resulted in both a higher Cotton effect
type of spectrum with a positive Cotton band at 270 nm of
than that produced by 1 mM Mg2⫹ and the formation of two
higher amplitude than that of the DNAzyme strand (Fig. 6C).
characteristic, discrete peaks (265 nm and 269 nm). At concen-
Both the lack of the symmetry for this band and the appearance
trations higher than 1 mM Mg2⫹, the height of these peaks is
of the weaker negative effect at 244 nm and positive effect at
reversed and kept practically unchanged, even at higher Mg2⫹
223 nm are characteristic for the spectrum. Upon addition of an
concentrations of up to 25 mM. The positive CD band of the
initial amount of Mg2⫹ (0.5 mM), the amplitude of the positive
Mg2⫹-free DNAzyme, although rather broad (276 –278 nm), is
Cotton effect rises substantially, and the peak is shifted down
much more regular (Fig. 6B) than that of the 21-mer oligode-
to 268 nm. No further increase of the Cotton effect was ob-
oxynucleotide. The spectrum also contains a negative effect at
served above 1 mM Mg2⫹. The results presented above indicate
249 nm and a weak, positive effect at 221 nm. This result
that, upon binding of Mg2⫹, the global geometry of the
indicates that some secondary structure exists for the 35-mer
DNAzyme adopts a compact structure projecting flanking arms
Structural Transitions Induced in the DNAzyme by Mg2⫹
Kinetic parameters for binding of 3DE or antisense oligodeoxynucleotide 3(1245-1265) to
immobilized RNA substrate and their dependence upon Mg2⫹ concentration
The 3⬘ end biotinylated RNA substrate (GUUCCACUCGUUAUCUUC) was immobilized on a BIAcore™ CM5 sensor chip coated with avidin
(⌬RU ⫽ 5000). The analyte was injected at a flow of 5 l/min, and measurements were done at 37 °C using 50 mM Tris, pH 8.0, containing divalentcations at the indicated concentrations. MeO-3DE used in these experiments was inactive due to a single base substitution (G6 3 A). Theinteraction of the corresponding oligodeoxynucleotide 3(1245-1265) containing both flanking arms of the 3DE with the same RNA substrate wasanalyzed under the same conditions. This oligodeoxynucleotide was modified as in MeO-3DE. The k
were determined from the
association and dissociation phases, respectively, with four different concentrations of the DNAzyme and the oligodeoxynucleotide. Apparent KAcorresponds to k /k
ratio. Data are shown as a mean value of four separate analyses.
kon (1/Ms)
koff (1/s)
(8.23 ⫾ 0.53) ⫻ 104
(1.98 ⫾ 0.94) ⫻ 10⫺3
(2.25 ⫾ 0.43) ⫻ 107
(8.47 ⫾ 0.77) ⫻ 104
(2.99 ⫾ 1.05) ⫻ 10⫺3
(3.16 ⫾ 1.23) ⫻ 107
(1.23 ⫾ 0.28) ⫻ 105
(2.84 ⫾ 1.53) ⫻ 10⫺3
(6.91 ⫾ 2.60) ⫻ 107
1.20 ⫾ 0.11) ⫻ 105
3.22 ⫾ 0.94) ⫻ 10⫺3
2.74 ⫾ 0.84) ⫻ 107
9.51 ⫾ 0.15) ⫻ 104
3.09 ⫾ 0.86) ⫻ 10⫺3
3.05 ⫾ 1.01) ⫻ 107
8.55 ⫾ 0.79) ⫻ 104
2.84 ⫾ 0.94) ⫻ 10⫺3
5.71 ⫾ 2.71) ⫻ 107
(3.44 ⫾ 1.04) ⫻ 104
(1.41 ⫾ 0.49) ⫻ 10⫺4
(1.64 ⫾ 0.29) ⫻ 108
(6.04 ⫾ 2.10) ⫻ 104
(3.76 ⫾ 0.53) ⫻ 10⫺4
(1.78 ⫾ 0.88) ⫻ 108
(8.49 ⫾ 2.15) ⫻ 104
(5.90 ⫾ 0.18) ⫻ 10⫺4
(1.46 ⫾ 0.40) ⫻ 108
5.22 ⫾ 1.46) ⫻ 104
8.40 ⫾ 1.88) ⫻ 10⫺4
1.19 ⫾ 0.84) ⫻ 108
7.84 ⫾ 1.48) ⫻ 104
6.11 ⫾ 1.02) ⫻ 10⫺4
1.37 ⫾ 0.52) ⫻ 108
9.41 ⫾ 2.94) ⫻ 104
7.21 ⫾ 1.50) ⫻ 10⫺4
1.48 ⫾ 0.83) ⫻ 108
at right position and angles to bind substrate mRNA.
the divalent cation dependence and kinetic parameters of RNA
To further test this concept, the binding kinetics were di-
cleavage. Exhaustive studies on chimeric DNAzymes and sub-
rectly measured by surface plasmon resonance analysis in the
strates composed of DNA and RNA showed that both types of
presence of increasing concentrations of Mg2⫹. In these exper-
enzymes have a very similar catalytic mechanism (5). The
iments, biotinylated RNA substrate was attached to the avidin-
reactions have an identical dependence on pH, both demon-
coated sensor. To avoid cleavage of the RNA substrate, the
strate an inverse correlation between the pK of metal hydrates
inactive DNAzyme containing a single nucleotide substitution
and activity and solvent isotope effects, and thio effects on the
in the catalytic domain of 3DE was used. The antisense oli-
reactions are identical (24). The crystal structure of several
godeoxynucleotide 3(1245–1265) consisting of both flanking
unmodified and modified hammerhead RNA in the absence of
arms (and thus antisense to 3 integrin subunit mRNA) was
divalent metal ions has been solved (28, 30 –32). Cation binding
tested as a control. To clarify the effect of cations on association
sites and the mechanism by which they control enzymatic
and dissociation processes between the DNAzyme and its RNA
activity have been elucidated (30, 33). Five Mg2⫹ sites are seen
substrate, we determined the parameters of k , k , and K in
in the crystal structure of the Mg2⫹-soaked freeze-trapped con-
the binding reactions between RNA substrate and either
formational intermediate of the hammerhead ribozyme, which
DNAzyme or the antisense oligodeoxynucleotide (Table II). The
could be divided into two groups based on their roles in cata-
binding affinity of 3 DNAzyme to the RNA substrate was
lytic activity. The first group consists of Mg2⫹ sites that upon
dependent upon the concentration of Mg2⫹. The association
binding of cations induce folding of the enzyme into its active
constant determined in the presence of 15 mM Mg2⫹ was sig-
conformation. The second group includes sites occupied by
nificantly higher (p ⬍ 0.001) than that observed at 5 mM of
Mg2⫹ bound directly to the optically active oxygen showing R
Mg2⫹. However, it was still much lower than the K describing
diastereomeric form (pro-R) at the cleavage site. The hammer-
the interaction of the antisense oligodeoxynucleotide with the
head ribozyme can cleave its own RNA, and this activity re-
same RNA fragment. Interestingly, the binding affinity of the
quires one or more catalytic divalent metal ions, one of which
antisense oligodeoxynucleotide to the RNA substrate did not
ionizes the 2⬘-hydroxyl at the cleavage site. The newly gener-
depend upon cation concentration, and regardless of the Mg2⫹
ated nucleophile attacks the adjacent phosphate by an in-line
presence, it was almost an order of magnitude higher than that
mechanism. The same metal ion, or perhaps another, stabilizes
of the DNAzyme.
the pentacoordinated phosphate transition state by binding
directly to the pro-R phosphate oxygen. The reaction generates
The intracellular ability of various 10 –23 DNAzymes to in-
5⬘-hydroxyl and 2⬘,3⬘-cyclic phosphate termini at the cleavage
hibit expression of the targeted proteins was evidenced by
several in vitro and in vivo studies (27), indicating their poten-
The structural effect of Mg2⫹ is well established in the ham-
tial advantages as biocatalysts in oligonucleotide therapy. De-
merhead ribozyme (35). In the absence of divalent metal ions,
spite tremendous therapeutic potential, the ability of the
the hammerhead structure is extended, with a disordered core,
DNAzyme to influence biological processes has not been deter-
but upon addition of metal ions, folding occurs in two distinct
mined at the molecular level. Because of its high flexibility, the
steps. Both events are well described by two-state transitions
three-dimensional structure of the DNAzyme molecule is not
induced by the non-cooperative binding of Mg2⫹. One can as-
yet known. Therefore, even a basic knowledge about the mech-
sume that similarly to the hammerhead ribozyme, metal ions
anism by which Mg2⫹ and other divalent cations regulate en-
will induce the folding of the DNAzyme molecule into the
zymatic activity of the 10 –23 DNAzyme is at present specula-
geometry required to facilitate the pathway into the transition
tive. The hypothetical mechanism for catalysis of RNA cleavage
state and will also bind at a specific location(s), where they can
by the DNAzyme is essentially based on assumptions that it
participate directly in the chemistry of the cleavage reaction.
behaves similarly to the hammerhead ribozyme, which also is
Data presented in this report show that binding of Mg2⫹ to
active in the presence of various divalent metal cations (28).
the 10 –23 DNAzyme induces significant rearrangement of the
Despite different compositions, the 10 –23 DNAzyme and the
catalytic loop, which leads to optimal folding of the molecule.
hammerhead ribozyme show many common features, including
This folding may occur in several distinct stages. The first
Structural Transitions Induced in the DNAzyme by Mg2⫹
transition induced by 0.5 mM Mg2⫹ results in the formation of
ribozyme (29, 30, 38). Interestingly, the binding affinity of the
a compact structure of the DNAzyme. The DNAzyme in such a
DNAzyme to RNA increased linearly in the presence of divalent
state binds weakly to its RNA substrate and lacks catalytic
cations when they were used in the range from 0 to 15 mM, but
activity. In the next stage observed at concentrations up to 5
it was still much lower than that of the antisense oligode-
mM Mg2⫹, the flanking arms are projected into the proper
oxynucleotide consisting of the DNAzyme flanking arms. (v)
position to bind the RNA substrate. Under such conditions, the
Essentially the same effect on the folding, binding affinity to
DNAzyme binds efficiently to the substrate and shows substan-
RNA substrate and catalytic activity of the DNAzyme were
tial catalytic activity. Further increase of Mg2⫹ leads to the
found when other divalent cations such as Ca2⫹ and Mn2⫹ were
final transition, involving formation of the completely orga-
used in the same concentration range. Monovalent cations such
nized catalytic domain of the DNAzyme. Such a mechanism is
as Na⫹, K⫹, and Li⫹ added in place of Mg2⫹ even at much
supported by the following observations: (i) E
higher concentrations of up to 1 M did not show any effect.
-Rhod rapidly increases in the range from 0 to 5
Acknowledgments—We thank Dr. A. Okruszek for synthesis of oli-
M cations and reaches a plateau value by 5 mM Mg2⫹, indi-
cating at this concentration the shortest distance between the
energy donor and acceptor. In the absence of Mg2⫹, theDNAzyme is inactive and its catalytic core is essentially un-
folded. With the addition of 5 mM Mg2⫹, the orientation of the
1. Santoro, S. W., and Joyce, G. F. (1997) Proc. Natl. Acad. Sci. U. S. A. 94,
catalytic loop changes, and the distance between 5⬘ and 3⬘ ends
almost reaches the value characteristic for the DNAzyme in
2. Cieslak, M., Niewiarowska, J., Nawrot, M., Koziolkiewicz, M., Stec, W. J., and
Cierniewski, C. S. (2002) J. Biol. Chem. 277, 6779 – 6787
complex with its RNA substrate. Under these conditions, the
3. Liu, C., Cheng, R., Sun, L. Q., and Tien, P. (2001) Biochem. Biophys. Res.
DNAzyme shows substantial enzymatic activity. A significant
Commun. 284, 1077–1082
4. Zhang, L., Gasper, W. J., Stass, S. A., Ioffe, O. B., Davis, M. A., and Mixson,
increase in the catalytic activity of the DNAzyme is observed
A. J. (2002) Cancer Res. 62, 5463–5469
when the Mg2⫹ concentration is increased from 5 to 15 mM,
5. He, Q. C., Zhou, J. M., Zhou, D. M., Nakamatsu, Y., Baba, T., and Taira, K.
suggesting that additional structural alterations within the
(2002) Biomacromolecules 3, 69 – 83
6. Santoro, S. W., and Joyce, G. F. (1998) Biochemistry 37, 13330 –13342
catalytic loop have occurred, even though there was no further
7. Chiu, T. K., and Dickerson, R. E. (2000) J. Mol. Biol. 25, 915–945
change in E
. The hyperbolic concentration dependence of
8. Welche, J. B., Duckett, D. R., and Lilley, D. M. (1993) Nucleic Acids Res. 21,
the end-to-end distance with a midpoint of ⬃2 mM Mg2⫹, which
9. Soyfer, V. N., and Potaman, V. N. (1966) Triple-helical Nucleic Acids, p. 360,
is significantly lower than that characteristic for the chemical
Springer, New York
cleavage step (Fig. 2), indicates that structural changes in-
10. Brukner, I., Susic, S., Dlakic, M., Savic, A., and Pongor, S. (1994) J. Mol. Biol.
11, 26 –32
duced in the DNAzyme occur at much lower Mg2⫹ concentra-
11. Dahlgreen, P. R., and Lyubchenko, Y. L. (2002) Biochemistry 41, 11372–11378
tions than those required for the catalytic properties. This
12. Scott, W., and Klug, A. (1996) Trends Biochem. Sci. 21, 220 –224
13. Cierniewski, C. S., Babinska, A., Swiatkowska, M., Wilczynska, M., Okruszek,
conclusion is supported by the observation that the mutated
A., and Stec, W. (1995) Eur. J. Biochem. 227, 494 – 499
inactive variant of the MeO-3DE, with the shortened catalytic
14. Okumoto, Y., and Sugimoto, N. (2000) J. Inorg. Chem. 82, 189 –195
loop, adopts the compact structure at a much lower Mg2⫹
15. Jaffe, E. A., Minich, R., Adelman, B., Becker, C. G., and Nachman, R. L. (1976)
J. Exp. Med. 144, 209 –221
concentration, indicating that such a structural transition is
16. Jaffe, E. A., Nachman, R. L., Becher, C. G., and Minich, C. R. (1973) J. Clin.
not sufficient to gain catalytic activity. (ii) The fluorescence
Invest. 52, 2745–2756
17. Clegg, R. M. (1992) Methods Enzymol. 211, 353–388
anisotropy of Flu-MeO-3DE-Rhod doubles after exposure to
18. Clegg, R. M., Murchie, A. I. H., Zechel, A., and Lilley, D. M. J. (1993) (1993)
the increasing concentrations of Mg2⫹ or Ca2⫹ and reaches the
Proc. Natl. Acad. Sci. U. S. A. 90, 2994 –2998
maximum at their concentration of 1–5 m
19. Bondeson, K., Frostell-Karlsson, A., Fagerstam, L., and Magnusson, G. (1993)
M, indicating the
Anal. Biochem. 214, 245–251
increased condensation state of the molecule under these con-
20. Kumar, P. K. R., Zhou, D.-M., Yoshinari, K., and Taira, K. (1996) in Catalytic
ditions. (iii) Saturation effects of Mg2⫹ concentrations detected
RNA, Nucleic Acids and Molecular Biology (Eckstein, F., and Lilley,
by CD spectroscopy were produced in the range from 0.5 to 2.0
D. M. J., eds) Vol. 10, pp. 217–230, Springer-Verlag, Berlin
21. Dass, C. R., Saravolac, E. G., Li, Y., and Sun, L. Q. (2002) Antisense Nucleic
mM, i.e. somewhat lower than that described by other tech-
Acid Drug Dev. 12, 289 –299
niques. As expected, spectra of the DNA/RNA hybrid (positive
22. Zhou, D.-M., Usman, N., Wincott, F. E., Matulic-Adamic, J., Orita, M., Zhang,
L.-H., Komiyama, M., Kumar, P. K. R., and Taira, K. (1996) J. Am. Chem.
effect at 269 nm), used as a referenced structure close to a
Soc. 118, 5862–5866
regular A-type helix (36, 37), were much less sensitive to Mg2⫹
23. Lakowicz, J. R. (1983) Principles of Fluorescence Spectroscopy, Plenum Press,
than those of the DNAzyme strand and its complex with RNA.
24. Ota, N., Warashina, M., Hirano, K., Hatanaka, K., and Taira, K. (1998) Nucleic
The Mg2⫹-free MeO-3DE strand (effect at 276 –278 nm) un-
Acids Res. 26, 3385–3391
dergoes a considerable change both upon Mg2⫹ binding (276
25. Clark, C. L., Cecil, P. K., Singh, D., and Gray, D. M. (1997) Nucleic Acids Res.
25, 4098 – 4105
nm) and further structural stabilization upon binding to the
26. Zuker, M. (2003) Nucleic Acids Res. 31, 1–10
target RNA strand. It should be emphasized that these confor-
27. Khachigian, L. M. (2002) Curr. Opin. Mol. Ther. 4, 119 –121
mational changes take place at a Mg2⫹ concentration as low as
28. Dahm, S. C., and Uhlenback, O. C. (1991) Biochemistry 30, 9464 –9469
29. Pley, H. W., Flaherty, K. M., and McKay, D. B. (1994) Nature, 372, 68 –74
0.5 mM. An overall similarity of the DNAzyme/RNA complex
30. Pley, H. W., Flaherty, K. M., and McKay, D. B. (1994) Nature 372, 111–113
spectra (Fig. 6C) to that typical for A-type RNA/DNA hybrids
31. Scott, W. G., Finch, J. T., and Klug, A. (1995) Cell 81, 991–1002
32. Ruffner, D. E., Stormo, G. D., and Uhlenbeck, O. C. (1990) Biochemistry 29,
was observed (25). This finding also confirms an earlier obser-
vation based upon various Mg2⫹-free DNAzyme complexes that
33. McKay, D. B. (1996) RNA 2, 395– 403
the hybrid nature of the flanking arms strongly influences their
34. Scott, W. G., Murray, J. B., Arnold, J. R. P., Stoddard, B. L., and Klug, A.
(1996) Science 274, 2065–2069
global geometry (24). (iv) The final form adopted at 15 mM
35. Bassi, G. S., Murchie, A. I., Walter, F., Clegg, R. M., and Lilley, D. M. (1997)
Mg2⫹, showing high binding affinity toward the mRNA sub-
EMBO J. 16, 7481–7489
36. Moore, D. S., and Wagner, T. E. (1974) Biopolymers, 13, 977–986
strate, is in good agreement with the global form of the struc-
37. Fairall, L., Martin, S., and Rhodes, D. (1989) EMBO J., 8, 1809 –1817
ture observed under the same conditions for the hammerhead
38. Breaker, R. R., and Joyce, G. F. (1995) Chem. Biol. 2, 655– 660
Source: http://www.ibch.poznan.pl/adamiak/pub/47987.pdf
A Provider's Guide for the Care of Women with Physical Disabilities and Chronic Suzanne C. Smeltzer, RN, EdD, FAAN Professor & Director, Nursing Research Director, Health Promotion for Women with Disabilities Project Villanova University College of Nursing 800 Lancaster Avenue Villanova, PA 19085 Phone: 610-519-6828 Fax: 610-519-7650 Nancy C. Sharts-Hopko, RN, PhD, FAAN
Case report of oral melanoma in a king penguin (Aptenodytes patagonicus) found by cooperation between Kosei Inui1, Miyuki Hamaguchi1, Satomi Yonezawa1, Makiko Nishizawa1, Takeshi Wada1, Kazutoshi Takami2 1: Naniwa-Hone-Hone-Dan, Osaka Museum of Natural History 2: Osaka Municipal Tennoji Zoological Gardens AZEC 2013, Fukuoka King penguin (Aptenodytes patagonicus)





